Phagosome
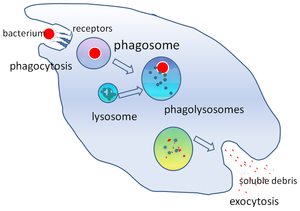
In cell biology, a phagosome is a vesicle formed around a particle engulfed by a phagocyte via phagocytosis. Professional phagocytes include macrophages, neutrophils, and dendritic cells (DCs).[1]
A phagosome is formed by the fusion of the cell membrane around a microorganism, a senescent cell or an apoptotic cell. Phagosomes have membrane-bound proteins to recruit and fuse with lysosomes to form mature phagolysosomes. The lysosomes contain hydrolytic enzymes and reactive oxygen species (ROS) which kill and digest the pathogens. Phagosomes can also form in non-professional phagocytes, but they can only engulf a smaller range of particles, and do not contain ROS. The useful materials (e.g. amino acids) from the digested particles are moved into the cytosol, and waste is removed by exocytosis. Phagosome formation is crucial for tissue homeostasis and both innate and adaptive host defense against pathogens.
However, some bacteria can exploit phagocytosis as an invasion strategy. They either reproduce inside of the phagolysosome (e.g. Coxiella spp.)[2] or escape into the cytoplasm before the phagosome fuses with the lysosome (e.g. Rickettsia spp.).[3] Many Mycobacteria, including Mycobacterium tuberculosis[4][5] and Mycobacterium avium paratuberculosis,[6] can manipulate the host macrophage to prevent lysosomes from fusing with phagosomes and creating mature phagolysosomes. Such incomplete maturation of the phagosome maintains an environment favorable to the pathogens inside it.[7]
Formation
[edit]Phagosomes are large enough to degrade whole bacteria, or apoptotic and senescent cells, which are usually >0.5μm in diameter.[8] This means a phagosome is several orders of magnitude bigger than an endosome, which is measured in nanometres.
Phagosomes are formed when pathogens or opsonins bind to a transmembrane receptor, which are randomly distributed on the phagocyte cell surface. Upon binding, "outside-in" signalling triggers actin polymerisation and pseudopodia formation, which surrounds and fuses behind the microorganism. Protein kinase C, phosphoinositide 3-kinase, and phospholipase C (PLC) are all needed for signalling and controlling particle internalisation.[9] More cell surface receptors can bind to the particle in a zipper-like mechanism as the pathogen is surrounded, increasing the binding avidity.[10] Fc receptor (FcR), complement receptors (CR), mannose receptor and dectin-1 are phagocytic receptors, which means that they can induce phagocytosis if they are expressed in non-phagocytic cells such as fibroblasts.[11] Other proteins such as Toll-like receptors are involved in pathogen pattern recognition and are often recruited to phagosomes but do not specifically trigger phagocytosis in non-phagocytic cells, so they are not considered phagocytic receptors.
Opsonisation
[edit]Opsonins are molecular tags such as antibodies and complements that attach to pathogens and up-regulate phagocytosis. Immunoglobulin G (IgG) is the major type of antibody present in the serum. It is part of the adaptive immune system, but it links to the innate response by recruiting macrophages to phagocytose pathogens. The antibody binds to microbes with the variable Fab domain, and the Fc domain binds to Fc receptors (FcR) to induce phagocytosis.
Complement-mediated internalisation has much less significant membrane protrusions, but the downstream signalling of both pathways converge to activate Rho GTPases.[12] They control actin polymerisation which is required for the phagosome to fuse with endosomes and lysosomes.
Non-phagocytic cells
[edit]Other non-professional phagocytes have some degree of phagocytic activity, such as thyroid and bladder epithelial cells that can engulf erythrocytes and retinal epithelial cells that internalise retinal rods.[8] However non-professional phagocytes do not express specific phagocytic receptors such as FcR and have a much lower rate of internalisation.
Some invasive bacteria can also induce phagocytosis in non-phagocytic cells to mediate host uptake. For example, Shigella can secrete toxins that alter the host cytoskeleton and enter the basolateral side of enterocytes.[13]
Structure
[edit]As the membrane of the phagosome is formed by the fusion of the plasma membrane, the basic composition of the phospholipid bilayer is the same. Endosomes and lysosomes then fuse with the phagosome to contribute to the membrane, especially when the engulfed particle is very big, such as a parasite.[14] They also deliver various membrane proteins to the phagosome and modify the organelle structure.
Phagosomes can engulf artificial low-density latex beads and then purified along a sucrose concentration gradient, allowing the structure and composition to be studied.[15] By purifying phagosomes at different time points, the maturation process can also be characterised. Early phagosomes are characterised by Rab5, which transition into Rab7 as the vesicle matures into late phagosomes.
Maturation process
[edit]The nascent phagosome is not inherently bactericidal. As it matures, it becomes more acidic from pH 6.5 to pH 4, and gains characteristic protein markers and hydrolytic enzymes. The different enzymes function at various optimal pH, forming a range so they each work in narrow stages of the maturation process. Enzyme activity can be fine-tuned by modifying the pH level, allowing for greater flexibility. The phagosome moves along microtubules of the cytoskeleton, fusing with endosomes and lysosomes sequentially in a dynamic "kiss-and-run" manner.[16] This intracellular transport depends on the size of the phagosomes. Larger organelles (with a diameter of about 3 μm) are transported very persistently from the cell periphery towards the perinuclear region whereas smaller organelles (with a diameter of about 1 μm) are transported more bidirectionally back and forth between cell center and cell periphery.[17] Vacuolar proton pumps (v-ATPase) are delivered to the phagosome to acidify the organelle compartment, creating a more hostile environment for pathogens and facilitating protein degradation. The bacterial proteins are denatured in low pH and become more accessible to the proteases, which are unaffected by the acidic environment. The enzymes are later recycled from the phagolysosome before egestion so they are not wasted. The composition of the phospholipid membrane also changes as the phagosome matures.[15]
Fusion may take minutes to hours depending on the contents of the phagosome; FcR or mannose receptor-mediated fusion last less than 30 minutes, but phagosomes containing latex beads may take several hours to fuse with lysosomes.[8] It is suggested that the composition of the phagosome membrane affects the rate of maturation. Mycobacterium tuberculosis has a very hydrophobic cell wall, which is hypothesised to prevent membrane recycling and recruitment of fusion factors, so the phagosome does not fuse with lysosomes and the bacterium avoids degradation.[18]
Smaller lumenal molecules are transferred by fusion faster than larger molecules, which suggests that a small aqueous channel forms between the phagosome and other vesicles during "kiss-and-run", through which only limited exchange is allowed.[8]
Fusion regulation
[edit]Shortly after internalisation, F-actin depolymerises from the newly formed phagosome so it becomes accessible to endosomes for fusion and delivery of proteins.[8] The maturation process is divided into early and late stages depending on characteristic protein markers, regulated by small Rab GTPases. Rab5 is present on early phagosomes, and controls the transition to late phagosomes marked by Rab7.[19]
Rab5 recruits PI-3 kinase and other tethering proteins such as Vps34 to the phagosome membrane, so endosomes can deliver proteins to the phagosome. Rab5 is partially involved in the transition to Rab7, via the CORVET complex and the HOPS complex in yeast.[19] The exact maturation pathway in mammals is not well understood, but it is suggested that HOPS can bind Rab7 and displace the guanosine nucleotide dissociation inhibitor (GDI).[20] Rab11 is involved in membrane recycling.[21]
Phagolysosome
[edit]The phagosome fuses with lysosomes to form a phagolysosome, which has various bactericidal properties. The phagolysosome contains reactive oxygen and nitrogen species (ROS and RNS) and hydrolytic enzymes. The compartment is also acidic due to proton pumps (v-ATPases) that transport H+ across the membrane, used to denature the bacterial proteins.
The exact properties of phagolysosomes vary depending on the type of phagocyte. Those in dendritic cells have weaker bactericidal properties than those in macrophages and neutrophils. Also, macrophages are divided into pro-inflammatory "killer" M1 and "repair" M2. The phagolysosomes of M1 can metabolise arginine into highly reactive nitric oxide, while M2 use arginine to produce ornithine to promote cell proliferation and tissue repair.[22]
Function
[edit]Pathogen degradation
[edit]Macrophages and neutrophils are professional phagocytes in charge of most of the pathogen degradation, but they have different bactericidal methods. Neutrophils have granules that fuse with the phagosome. The granules contain NADPH oxidase and myeloperoxidase, which produce toxic oxygen and chlorine derivatives to kill pathogens in an oxidative burst. Proteases and anti-microbial peptides are also released into the phagolysosome. Macrophages lack granules, and rely more on phagolysosome acidification, glycosidases, and proteases to digest microbes.[21] Phagosomes in dendritic cells are less acidic and have much weaker hydrolytic activity, due to a lower concentration of lysosomal proteases and even the presence of protease inhibitors.
Inflammation
[edit]Phagosome formation is tied to inflammation via common signalling molecules. PI-3 kinase and PLC are involved in both the internalisation mechanism and triggering inflammation.[9] The two proteins, along with Rho GTPases, are important components of the innate immune response, inducing cytokine production and activating the MAP kinase signalling cascade. Pro-inflammatory cytokines including IL-1β, IL-6, TNFα, and IL-12 are all produced.[8]
The process is tightly regulated and the inflammatory response varies depending on the particle type within the phagosome. Pathogen-infected apoptotic cells will trigger inflammation, but damaged cells that are degraded as part of the normal tissue turnover do not. The response also differs according to the opsonin-mediated phagocytosis. FcR and mannose receptor-mediated reactions produce pro-inflammatory reactive oxygen species and arachidonic acid molecules, but CR-mediated reactions do not result in those products.[8]
Antigen presentation
[edit]Immature dendritic cells (DCs) can phagocytose, but mature DCs cannot due to changes in Rho GTPases involved in cytoskeleton remodelling.[21] The phagosomes of DCs are less hydrolytic and acidic than those of macrophages and neutrophils, as DCs are mainly involved in antigen presentation rather than pathogen degradation. They need to retain protein fragments of a suitable size for specific bacterial recognition, so the peptides are only partially degraded.[21] Peptides from the bacteria are trafficked to the Major Histocompatibility Complex (MHC). The peptide antigens are presented to lymphocytes, where they bind to T-cell receptors and activates T-cells, bridging the gap between innate and adaptive immunity.[9] This is specific to mammals, birds, and jawed fish, as insects do not have adaptive immunity.[23]

Nutrient
[edit]Ancient single-celled organisms such as amoeba use phagocytosis as a way to acquire nutrients, rather than an immune strategy. They engulf other smaller microbes and digest them within the phagosome of around one bacterium per minute, which is much faster than professional phagocytes.[24] For the soil amoeba Dictyostelium discoideum, their main food source is the bacteria Legionella pneumophila, which causes Legionnaire's disease in humans.[25] Phagosome maturation in amoeba is very similar to that in macrophages, so they are used as a model organism to study the process.[16]
Tissue clearance
[edit]Phagosomes degrade senescent cells and apoptotic cells to maintain tissue homeostasis. Erythrocytes have one of the highest turnover rates in the body, and they are phagocytosed by macrophages in the liver and spleen. In the embryo, the process of removing dead cells is not well-characterised, but it is not performed by macrophages or other cells derived from hematopoietic stem cells.[26] It is only in the adult that apoptotic cells are phagocytosed by professional phagocytes. Inflammation is only triggered by certain pathogen- or damage-associated molecular patterns (PAMPs or DAMPs), the removal of senescent cells is non-inflammatory.[14]
Autophagosome
[edit]Autophagosomes are different from phagosomes in that they are mainly used to selectively degrade damaged cytosolic organelles such as mitochondria (mitophagy). However, when the cell is starved or stressed, autophagosomes can also non-selectively degrade organelles to provide the cell with amino acids and other nutrients.[27] Autophagy is not limited to professional phagocytes, it is first discovered in rat hepatocytes by cell biologist Christian de Duve.[28] Autophagosomes have a double membrane, the inner one from the engulfed organelle, and the outer membrane is speculated to be formed from the endoplasmic reticulum or the ER-Golgi Intermediate Compartment (ERGIC).[29] The autophagosome also fuses with lysosomes to degrade its contents. When M. tuberculosis inhibit phagosome acidification, Interferon gamma can induce autophagy and rescue the maturation process.[30]
Bacterial evasion and manipulation
[edit]Many bacteria have evolved to evade the bactericidal properties of phagosomes or even exploit phagocytosis as an invasion strategy.
- Mycobacterium tuberculosis target M2 macrophages at the lower parts of the respiratory pathway, which do not produce ROS.[31] M. tuberculosis can also manipulate the signalling pathways by secreting phosphatases such as PtpA and SapM, which disrupt protein recruitment and block phagosome acidification.[8][32]
- Legionella pneumophila can re-model the phagosome membrane to imitate vesicles in other parts of the secretory pathway, so lysosomes do not recognise the phagosome and do not fuse with it. The bacterium secretes toxins that interfere with host trafficking, so the Legionella-containing vacuole recruits membrane proteins usually found on the endoplasmic reticulum or the ERGIC.[33] This re-directs secretory vesicles to the modified phagosome and deliver nutrients to the bacterium.
- Listeria monocytogenes secretes a pore-forming protein listeriolysin O so the bacterium can escape the phagosome into the cytosol. Listeriolysin is activated by the acidic environment of the phagosome.[34] In addition, Listeria secrete two phospholipase C enzymes that facilitate in phagosome escape.
See also
[edit]References
[edit]- ^ Robinson & Babcock 1998, p. 187 and Ernst & Stendahl 2006, pp. 7–10
- ^ Hackstadt T, Williams JC (May 1981). "Biochemical stratagem for obligate parasitism of eukaryotic cells by Coxiella burnetii". Proceedings of the National Academy of Sciences of the United States of America. 78 (5): 3240–4. doi:10.1073/pnas.78.5.3240. PMC 319537. PMID 6942430.
- ^ Winkler HH (1990). "Rickettsia Species (As Organisms)". Annual Review of Microbiology. 44: 131–153. doi:10.1146/annurev.micro.44.1.131. PMID 2252380.
- ^ MacMicking JD, Taylor GA, McKinney JD (October 2003). "Immune control of tuberculosis by IFN-gamma-inducible LRG-47". Science. 302 (5645): 654–9. Bibcode:2003Sci...302..654M. doi:10.1126/science.1088063. PMID 14576437. S2CID 83944695.
- ^ Vandal OH, Pierini LM, Schnappinger D, Nathan CF, Ehrt S (August 2008). "A membrane protein preserves intrabacterial pH in intraphagosomal Mycobacterium tuberculosis". Nature Medicine. 14 (8): 849–54. doi:10.1038/nm.1795. PMC 2538620. PMID 18641659.
- ^ Kuehnel MP, Goethe R, Habermann A, Mueller E, Rohde M, Griffiths G, Valentin-Weigand P (August 2001). "Characterization of the intracellular survival of Mycobacterium avium ssp. paratuberculosis: phagosomal pH and fusogenicity in J774 macrophages compared with other mycobacteria". Cellular Microbiology. 3 (8): 551–66. doi:10.1046/j.1462-5822.2001.00139.x. PMID 11488816. S2CID 8962102.
- ^ Tessema MZ, Koets AP, Rutten VP, Gruys E (November 2001). "How does Mycobacterium avium subsp. paratuberculosis resist intracellular degradation?". The Veterinary Quarterly. 23 (4): 153–62. doi:10.1080/01652176.2001.9695105. PMID 11765232.
- ^ a b c d e f g h Aderem A, Underhill DM (April 1999). "Mechanisms of phagocytosis in macrophages". Annual Review of Immunology. 17 (1): 593–623. doi:10.1146/annurev.immunol.17.1.593. PMID 10358769.
- ^ a b c Aderem A (June 2003). "Phagocytosis and the inflammatory response". The Journal of Infectious Diseases. 187 Suppl 2 (s2): S340-5. doi:10.1086/374747. PMID 12792849.
- ^ Dupuy AG, Caron E (June 2008). "Integrin-dependent phagocytosis: spreading from microadhesion to new concepts". Journal of Cell Science. 121 (11): 1773–83. doi:10.1242/jcs.018036. PMID 18492791.
- ^ Underhill DM, Ozinsky A (April 2002). "Phagocytosis of microbes: complexity in action". Annual Review of Immunology. 20 (1): 825–52. doi:10.1146/annurev.immunol.20.103001.114744. PMID 11861619.
- ^ Kaplan G (August 1977). "Differences in the mode of phagocytosis with Fc and C3 receptors in macrophages". Scandinavian Journal of Immunology. 6 (8): 797–807. doi:10.1111/j.1365-3083.1977.tb02153.x. PMID 561436. S2CID 25373560.
- ^ Köhler H, Rodrigues SP, McCormick BA (March 2002). "Shigella flexneri Interactions with the Basolateral Membrane Domain of Polarized Model Intestinal Epithelium: Role of Lipopolysaccharide in Cell Invasion and in Activation of the Mitogen-Activated Protein Kinase ERK". Infection and Immunity. 70 (3): 1150–8. doi:10.1128/IAI.70.3.1150-1158.2002. PMC 127798. PMID 11854195.
- ^ a b Desjardins M, Houde M, Gagnon E (October 2005). "Phagocytosis: the convoluted way from nutrition to adaptive immunity". Immunological Reviews. 207 (1): 158–65. doi:10.1111/j.0105-2896.2005.00319.x. PMID 16181334. S2CID 30191200.
- ^ a b Desjardins M, Celis JE, van Meer G, Dieplinger H, Jahraus A, Griffiths G, Huber LA (December 1994). "Molecular characterization of phagosomes". The Journal of Biological Chemistry. 269 (51): 32194–200. doi:10.1016/S0021-9258(18)31620-X. PMID 7798218.
- ^ a b Gotthardt D, Warnatz HJ, Henschel O, Brückert F, Schleicher M, Soldati T (October 2002). "High-resolution dissection of phagosome maturation reveals distinct membrane trafficking phases". Molecular Biology of the Cell. 13 (10): 3508–20. doi:10.1091/mbc.E02-04-0206. PMC 129962. PMID 12388753.
- ^ Keller S, Berghoff K, Kress H (December 2017). "Phagosomal transport depends strongly on phagosome size". Scientific Reports. 7 (1): 17068. Bibcode:2017NatSR...717068K. doi:10.1038/s41598-017-17183-7. PMC 5719076. PMID 29213131.
- ^ de Chastellier C, Thilo L (September 1997). "Phagosome maturation and fusion with lysosomes in relation to surface property and size of the phagocytic particle". European Journal of Cell Biology. 74 (1): 49–62. PMID 9309390.
- ^ a b Fairn GD, Grinstein S (August 2012). "How nascent phagosomes mature to become phagolysosomes". Trends in Immunology. 33 (8): 397–405. doi:10.1016/j.it.2012.03.003. PMID 22560866.
- ^ Kinchen JM, Ravichandran KS (April 2010). "Identification of two evolutionarily conserved genes regulating processing of engulfed apoptotic cells". Nature. 464 (7289): 778–82. Bibcode:2010Natur.464..778K. doi:10.1038/nature08853. PMC 2901565. PMID 20305638.
- ^ a b c d Savina A, Amigorena S (October 2007). "Phagocytosis and antigen presentation in dendritic cells". Immunological Reviews. 219 (1): 143–56. doi:10.1111/j.1600-065X.2007.00552.x. PMID 17850487. S2CID 1267478.
- ^ Mills CD (5 May 2015). "Anatomy of a discovery: m1 and m2 macrophages". Frontiers in Immunology. 6: 212. doi:10.3389/fimmu.2015.00212. PMC 4419847. PMID 25999950.
- ^ Warr GW (1997). "The adaptive immune system of fish". Developments in Biological Standardization. 90: 15–21. PMID 9270830.
- ^ Cosson P, Soldati T (June 2008). "Eat, kill or die: when amoeba meets bacteria". Current Opinion in Microbiology. 11 (3): 271–6. doi:10.1016/j.mib.2008.05.005. PMID 18550419.
- ^ Solomon JM, Rupper A, Cardelli JA, Isberg RR (May 2000). "Intracellular growth of Legionella pneumophila in Dictyostelium discoideum, a system for genetic analysis of host-pathogen interactions". Infection and Immunity. 68 (5): 2939–47. doi:10.1128/iai.68.5.2939-2947.2000. PMC 97507. PMID 10768992.
- ^ Lichanska AM, Hume DA (June 2000). "Origins and functions of phagocytes in the embryo" (PDF). Experimental Hematology. 28 (6): 601–11. doi:10.1016/s0301-472x(00)00157-0. hdl:20.500.11820/b865effa-680c-4f60-905e-ba9137f57613. PMID 10880746. S2CID 13336661.
- ^ Ding WX, Yin XM (July 2012). "Mitophagy: mechanisms, pathophysiological roles, and analysis". Biological Chemistry. 393 (7): 547–64. doi:10.1515/hsz-2012-0119. PMC 3630798. PMID 22944659.
- ^ Castro-Obregon S (2010). "The Discovery of Lysosomes and Autophagy". Nature Education. 3 (9): 49.
- ^ Ge L, Schekman R (January 2014). "The ER-Golgi intermediate compartment feeds the phagophore membrane". Autophagy. 10 (1): 170–2. doi:10.4161/auto.26787. PMC 4389871. PMID 24220263.
- ^ Gutierrez MG, Master SS, Singh SB, Taylor GA, Colombo MI, Deretic V (December 2004). "Autophagy is a defense mechanism inhibiting BCG and Mycobacterium tuberculosis survival in infected macrophages". Cell. 119 (6): 753–66. doi:10.1016/j.cell.2004.11.038. PMID 15607973.
- ^ Cambier CJ, Takaki KK, Larson RP, Hernandez RE, Tobin DM, Urdahl KB, Cosma CL, Ramakrishnan L (January 2014). "Mycobacteria manipulate macrophage recruitment through coordinated use of membrane lipids". Nature. 505 (7482): 218–22. doi:10.1038/nature12799. PMC 3961847. PMID 24336213.
- ^ Wong D, Chao JD, Av-Gay Y (February 2013). "Mycobacterium tuberculosis-secreted phosphatases: from pathogenesis to targets for TB drug development". Trends in Microbiology. 21 (2): 100–9. doi:10.1016/j.tim.2012.09.002. PMID 23084287.
- ^ Roy CR, Kagan JC (1 January 2013). Evasion of Phagosome Lysosome Fusion and Establishment of a Replicative Organelle by the Intracellular Pathogen Legionella pneumophila. Landes Bioscience.
- ^ Portnoy DA, Auerbuch V, Glomski IJ (August 2002). "The cell biology of Listeria monocytogenes infection: the intersection of bacterial pathogenesis and cell-mediated immunity". The Journal of Cell Biology. 158 (3): 409–14. doi:10.1083/jcb.200205009. PMC 2173830. PMID 12163465.
